Location: Home >> Detail
TOTAL VIEWS
J Sustain Res. 2026;8(2):e260044. https://doi.org/10.20900/jsr20260044
Department of Computational Science (Physics Discipline), University of Fort Hare, Faculty of Science and Agriculture, Private Bag X1314, Alice 5700, South Africa
* Correspondence: KeChrist Obileke
Anaerobic digestion (AD) converts organic wastes into biogas through sequential microbial processes whose rates depend on substrate characteristics, environmental conditions, and reactor configuration. Therefore, the aim of this review is to examine the application of kinetic modelling in relation to anaerobic digestion for biogas production. The review found that the first-order modified Gompertz, logistic, and transfer function models are the most employed for biogas production. Findings revealed that the modified Gompertz and logistic function models fit the mesophilic anaerobic digestion data well, whereas the transference model provides a better fit for the thermophilic anaerobic digestion data. Most studies have reported a robust correlation between experimental and model parameters, particularly in modelling biogas production from different waste sources. However, one main possible area of future study focuses on the scaling processes of kinetic modelling. The review presents a sensitivity analysis and trade-offs for model selection in kinetic modelling of anaerobic digestion for biogas production, as is common in most review papers.
Biogas is produced during the biological breakdown of organic matter in the absence of oxygen. In addition, it is the product formed during the anaerobic digestion (AD) process, which consists of the following compounds: CO2, CH4, H2S, H2, H2O, and other substances, depending on the composition of the substrate [1]. Subsequently, biogas consists of methane (CH4) and carbon dioxide (CO2), both greenhouse gases that are harmful to the environment; therefore, it is crucial that biogas undergo combustion before release into the atmosphere [2]. Anaerobic digestion is a process in which microorganisms convert organic matter into methane, carbon dioxide, and inorganic nutrients in the absence of oxygen [3]. There are four stages of AD: hydrolysis, acidogenesis, acetogenesis, and methanogenesis [4]. Hydrolysis, the first stage, is an enzyme-mediated process that converts insoluble organic compounds, such as proteins, fats, lipids, and carbohydrates, into soluble organic components, including amino acids, fatty acids, and monosaccharides [5]. It is achieved by solubilising and degrading particulate organic biopolymers and colloidal waste into soluble monomeric compounds [6]. The hydrolysis equation is given by [7]:
Acidogenesis is the second stage, during which the organic acid produced in hydrolysis is transformed into acetic acid, carbon dioxide, hydrogen gas, and acid derivatives [8]. It involves the production of an excessive concentration of hydrogen by an acidogenic bacterium and the degradation of organic matter to generate biogas, which is facilitated by different groups of bacteria. The main ones are acid-producing bacteria, known as acidogens, and methane-producing bacteria, referred to as methanogens [9]. At this stage, fatty acids are broken down into simpler components, including acetate and hydrogen, while volatile fatty acids are produced [10]. The equations are given by [11]
Acetogenesis is the third stage, in which compounds such as acetate, formaldehyde, hydrogen, and carbon dioxide are converted into methane and water. The simple molecules produced by acidogenesis are further digested by an acetogen bacterium [12]. The equations are given by [13]:
Methanogenesis, the final stage, is the process by which methanogens produce methane from acetic acid, water, and carbon dioxide [14]. The primary route is the fermentation of the major product of the acid-forming phase, acetic acid, to methane and carbon dioxide [15]. The equation for methanogenesis is given by [16]:
AD (anaerobic digestion) is a promising method for biogas production, in which organic matter is converted into biogas through the sequential involvement of different groups of bacteria [17]. The advantages of AD include relatively low operating costs, low energy consumption, applicability on both small and large scales, low sludge generation, and nutrient and chemical requirements [18].
Feedstocks are crucial for anaerobic digestion (AD) due to their high availability and the large quantities produced. They provide a consistent and rich source of organic matter for biogas production, offering a substantial, steady supply to enhance biogas yields [19]. However, the rapid breakdown of readily degradable materials can result in excessive accumulation of volatile fatty acids (VFAs), leading to acidification and limiting improvements in organic loading rates and biogas production in mono-digestion systems [20]. Therefore, maintaining a balanced nutrient balance within the system is critical to prevent VFA overproduction and ensure smooth biogas production. Many feedstocks are readily available globally as organic waste [21]. Co-digestion of different wastes offers a practical solution by combining their strengths and leveraging the advantages of diverse substrates, thereby improving the C/N ratio, reducing ammonia inhibition from nitrogen-rich materials, and mitigating acidification risks from VFA accumulation [22]. Integrating multiple feed stocks into the co-digestion process is thus a viable and economical option for enhanced biogas production and energy recovery [23]. Despite the advantages and potential of anaerobic digestion, kinetic modeling plays a role in advancing the technology.
Kinetic modelling refers to the mathematical techniques used to simulate and predict the rate of change in systems [24]. In anaerobic digestion, it quantifies the complex microbial breakdown of various organic wastes into biogas [25]. This technique also predicts and optimizes the biodegradation process to ensure methane yield is maximised and its process efficiency [26]. The use of mathematical models simulates the rate of biogas production from various feedstocks. This approach is vital for understanding the impact of feedstock composition and operating conditions on biogas yield and for scaling up AD technology to meet global energy and waste challenges. The kinetic model is said to predict the profile of gas composition and temperature inside the biogas digester as well as the overall performance for a given operating condition. Its parameters include biogas potential, maximum biogas rate, lag phase, and the coefficient of determination, all of which are simplified to facilitate fitting of experimental biogas production data. These examples enhance the efficiency and sustainability of the system design, as noted by [27], by providing valuable insights into the process. However, the conventional model or traditional method, such as the Anaerobic Digestion Model No. 1 (ADM1), offers a foundational framework but is constrained by computational complexity and requires precise calibration, which is a factor against scalability in large-scale applications. Although it is fundamental for simulating biogas production and understanding biological mechanisms, they struggle to accurately represent complex real-world systems [28]. To address these limitations, kinetic modelling offers a significant advantage by providing a detailed, time-dependent, mechanistic understanding of reactions. In addition, it allows the prediction of system behaviour over time under varied, complex, and non-ideal conditions. The key kinetic models include the modified Gompertz, first-order, cone, and logistic functions, among others. The advantages of kinetic modelling, as discussed relative to the traditional method and conventional models, motivate the review.
Research aim:
The aim of this study is to conduct an in-depth, comprehensive review of the application of kinetic modelling to anaerobic digestion and biogas production.
Research question:
●
●
●
●
●
Previous studies on the topic focus on a specific substrate or waste and are mostly published as original research articles. Furthermore, studies have been conducted on mathematical modelling to predict the behaviour of biogas digester systems using mathematical language. The present study differs from the previous studies as it is a review paper focusing on summarizing and carrying out an in-depth and comprehensive analysis of the kinetic modelling of different substrates for biogas production from existing published research in the absence of new experimental data. Also, the present study is a kinetic study of the rate of a chemical reaction and the equations involved through a mechanism. The major contributions of the review are as follows:
●
●
Databases such as ScienceDirect, Scopus, and Web of Science were used in the study, similar to [29]. Keywords such as biogas yield, prediction performance, reactor design, methane production, and renewable energy. Importantly, these keywords were used by the authors to identify relevant literature from peer-reviewed journals. It is worth noting that this was not constrained by the search time. Regarding the year of journal publications focusing on the study and its use, no limit was applied. However, all literature on the kinetic modelling of anaerobic digestion for biogas production published up to December 2025 was considered. The review considered original research, review articles, and book chapters on result search, including publications with selected keywords (topic, abstract, contributor, and keywords). The review applied both the inclusion and exclusion criteria as standard requirements in the design of a high-quality study. Inclusion criteria are the features the authors used in the study, whereas exclusion criteria are the features that were not included and thereby interfered with the study's success [30]. For inclusion criteria, the following were applied: publication in the form of original articles, review papers and book chapters from peer-reviewed journals. The review was written in English only. The type of publication employed in the study is a review article (narrative). On the contrary, examples of exclusion criteria include publications other than original articles, review papers from peer-reviewed journals, and languages other than English [29].
Different Wastes for Biogas ProductionBiogas production utilizes various organic waste streams, including agricultural waste (crop residues, manure), municipal waste (household food waste, yard trimmings), food waste (restaurant scraps, food processing waste), sewage waste (wastewater treatment sludge), and industrial waste (brewery, dairy, and meat processing waste) [31]. These waste types vary in composition, moisture content, and biodegradability, which in turn influence biogas yield and quality [32]. Agricultural and food waste tend to have high biodegradability and methane potential, whereas municipal and sewage waste may require pre-treatment to optimize biogas production [33]. Industrial waste, such as brewery waste, can be particularly suitable for biogas production due to its high organic content and consistency [34]. Utilizing these diverse waste streams for biogas production reduces waste disposal issues, greenhouse gas emissions, and reliance on fossil fuels, promoting a circular economy and sustainable energy generation [35].
Kinetic Models for Biogas ProductionThe First-Order Kinetic Model: This describes the rate of substrate degradation or product formation as proportional to the substrate concentration. The model is commonly used to model biogas production in anaerobic digestion systems, where the rate of biogas production is directly related to the concentration of organic matter being degraded. The model's simplicity and effectiveness make it a popular choice for predicting biogas yields and optimizing anaerobic digestion processes. By applying the First-Order Kinetic Model, researchers and practitioners can gain insights into the kinetics of biogas production [36]. The equation of the First Order Kinetic Model is given by:
where: y is the maximum biogas yield at digestion time t (mL/g); P is the maximum biogas production (mL/g); K is the rate constant; e is the 2.718282 (logarithmic constant); and t is the hydraulic retention time (days).
The Modified Gompertz Kinetic Model: A variation of the Gompertz model, often used to describe cumulative biogas production over time in anaerobic digestion systems. The Modified Gompertz Model is useful for predicting and optimizing biogas production, allowing researchers and practitioners to better understand the kinetics of anaerobic digestion processes and improve system design and operation. The Modified Gompertz kinetic model is given by Equation (6) [37].
where: y is the maximum biogas yield at digestion time t (mL/g); P is the maximum biogas production (mL/g); R_m-Biogas production rate (per day); λ is the Lag phase (day); e is 2.718282 (logarithmic constant), and t is the hydraulic retention time (day).
The Logistic Kinetic Model: This is a sigmoidal (S-shaped) curve, representing growth or product formation over time. This model is useful for modelling biogas production, microbial growth, and substrate degradation in anaerobic digestion systems. This model enables researchers and practitioners to understand and predict biogas production kinetics, facilitating the optimization of anaerobic digestion processes. Hence, it aligns with the global trend in kinetic analysis related to biogas production. The Logistic kinetic model holds that the biogas production rate is directly proportional to the amount of biogas already produced, the maximum production rate, and the maximum biogas production capacity. One application of such a model is the estimation of methane production in landfill leachate [38]. This model is expressed as given in Opurum (2021) [39] study as;
where: y represents the maximum biogas yield at digestion time t (mL/g); P is the maximum biogas production (mL/g); R_m is the biogas production rate (per day); λ is the Lag phase (day); e is the 2.718282 (logarithmic constant) and t is the hydraulic retention time (day).
The transference model: This is typically included to mathematically fit both inputs and outcomes in either black-box or curve-type models and is usually employed to evaluate the success rate of pretreatments [40]. The transference model originated from the first-order kinetic model, in which the kinetic constant is replaced by the ratio of the maximum methane production to the substrate's methane production potential [41]. The model is applicable for control purposes, for the conceptualization of a system, and for the lag phase in biogas production and biodegradable substrates [42,43]. The model is presented in Equation (8)
where: y is the maximum biogas yield at digestion time t (mL/g); P is the maximum Biogas production (mL/g); Rm is the biogas production rate (per day); λ is the Lag phase (day); e is the 2.718282 (logarithmic constant) and t represents the hydraulic retention time (day).
Cone Model: This type of kinetic model is an empirical model developed to evaluate methane production from a specific substrate in the presence of ruminal microorganisms [44]. It concerns model-predicted biogas production in anaerobic digestion systems, represented by an equation describing cumulative biogas production over time. Typically following a sigmoidal curve, the exact formula can vary depending on the specific application or modification of the Cone Model; it generally accounts for key parameters such as biogas production rate and lag phase [45]. The Cone Model is valuable for understanding and optimizing biogas production, enabling more efficient process management and system design in anaerobic digestion [46]. Similarly, it estimates methane production behaviour using the shape factor (Fs). Interestingly, the Fs indicate the presence or absence of the lag phase [47].
where BMP(t) (mL or mL/g VS) is the biogas or methane production at time (t). BMP(0) (mL or mL/g VS) is the maximum potential methane production, k (h–1 or d–1) is the rate constant, t (h or d) is the time, and n is dimensionless (sharpness or steepness of the curve).
Monod kinetics: This mathematical model describes the relationship between microbial growth rate and substrate concentration. This model is commonly applied in bioreactors, anaerobic digestion, and wastewater treatment [48]. The Monod equation describes a saturable relationship between microbial growth rate and substrate concentration, where growth rate increases with substrate concentration but eventually reaches a maximum [49]. The Monod model requires two key parameters: the maximum specific growth rate (μmax) and the half-saturation constant (Ks). These parameters help predict microbial growth rates and substrate consumption, making the Monod model a valuable tool for designing bioreactors and optimizing anaerobic digestion processes [50]. The Monod equation is given as expressed in Mamur et al. (2022) [51] as;
where: μ is the specific growth rate; μmax is the maximum specific growth rate; S is the concentration of the limiting substrate; Ks is the substrate affinity constant.
Anaerobic Digestion Model No. 1 (ADM1): This comprehensive kinetic model simulates anaerobic digestion processes and describes complex biochemical and physicochemical transformations [47]. Developed by the International Water Association, ADM1 encompasses multiple processes, including disintegration, hydrolysis, acidogenesis, acetogenesis, and methanogenesis, enabling the dynamic simulation of biogas production and system behaviour [52]. The model's versatility makes it valuable for designing, optimizing, and operating anaerobic digestion systems in various applications, including wastewater treatment and biogas production [53]. By accurately predicting system responses to different conditions, ADM1 supports process improvement and innovation in anaerobic digestion technology [54]. The main equation for ADM1 is a system of mass-balance differential equations for each component, such as the soluble (S) and particulate substrate (X), which takes the form [55].
where: Si: concentrate of the i-th soluble component, Xi: concentrate of the i-th particulate component, q: inflow and outflow rate, Vliq: liquid phase volume in the reactor, Si,in: concentrate of the i-th soluble component in the inflow, Si,liq: concentrate of the i-th soluble component in the liquid phase, vi,j: the stoichiometric coefficient for the i-th component in the j-th process, ρj: the kinetic rate of the j-th biochemical process.
Having looked at the various kinetic modelling techniques for anaerobic digestion to produce biogas, Table 1 briefly presents the strengths and weaknesses of these models.
Studies have used various kinetic models, including first-order, modified Gompertz, logistic, transfer, cone, and ADMI, across different waste types and conditions. The selected studies are presented in Table 2.
The primary application of kinetic modelling in biogas production involves optimising biogas digesters, predicting biogas yield, validating experimental data, and scaling up processes [9,120]. The kinetic models mentioned earlier have distinct features, strengths, and weaknesses. Nevertheless, the general application of these models focuses on simulation and improving the performance and design of biogas digesters [121]. Optimizing biogas digester processes involves controlling factors that affect biogas production, such as temperature, pH, and the carbon-to-nitrogen ratio. Maximising methane production rates, adjusting the substrate mixing ratio, and controlling the organic loading rate also help optimise biogas digester processes. According to Obileke et al. (2024) [122], optimizing process parameters can improve biogas digester performance, thereby increasing its production rate. Thus, it is also important in producing biogas from organic wastes [123]. Notably, biogas production can occur in either a batch or continuous system. The kinetic model equation, combined with temperature data and other parameters, is used to estimate biogas production, particularly in continuous systems. Due to the several experiments required and a significant amount of time and resources consumed in traditional methods. As a result, traditional methods become ineffective for optimization studies. Kinetic modelling in predicting biogas yield plays a crucial role in the growth of microorganisms, which is directly proportional to the rate of biogas production [124]. They are instrumental in predicting the biogas potential of any anaerobic digestion system [125]. Both the modified Gompertz model and the first-order kinetic model are widely used to predict biogas yield. Input parameters are crucial factors to consider in predicting biogas yield through kinetic analysis. For instance, the study conducted by [126] on the kinetic model of biogas yield from different vinasses. It was found that pH affected the modified Gompertz and first-order models. Therefore, the modified Gompertz model had the least variation from the measured biogas. Kinetic modelling in the validation of experimental data involves calibration by reproducing the experimental data. According to [127], mathematical modelling, including kinetic modelling, offers opportunities to simulate untested anaerobic digestion factors and validate experimental data. Bakraoui et al. (2020) [112] compared and adjusted experimental data using kinetic modelling with the modified Gompertz and first-order kinetic models. This implies that both models tend to produce methane and that a model could predict results. In this case, the modified Gompertz model showed the least deviation, ranging from 1% to 13%, compared with the first-order kinetic model, which showed deviations of 5% to 17%. Considering the application of kinetic modelling in scaling up processes, it has been applied in anaerobic digestion, ranging from lab-scale experiments to pilot and full-scale industrial reactors or plants [128]. This is specifically concerned with model parameters (lag phase and biogas potential) [127]. In this case, these model parameters are relevant for designing large-scale systems, thereby enabling the prediction and optimisation of system performance, including addressing the challenges [129].
Examples of Studies in Relation to the Application of Kinetic Modelling of Biogas ProductionKinetic modeling is applicable for the optimization of biogas digester, prediction of biogas yield, validation of experimental data and scaling processes. These are briefly discussed further.
Optimization of the biogas digesterA typical application of kinetic modelling to optimise the biogas digester process was reported by Bułkowska et al. (2025) [129]. Using the ADM1 mathematical model, the authors evaluated the conversion of organic compounds during the fermentation of maize silage. The study was conducted in a continuous-stirring anaerobic digester with a volume of 6 L, operating at 39 °C and a hydraulic retention time of 45 days. Parameters considered for the optimization include the disintegration constant (kdis) and hydrolysis concentration for carbohydrate (khyd_ch). The volatile fatty acid parameter includes the uptake constants for sugars, amino acids, higher fatty acids, and volatile fatty acids (Km and Ks). To identify numerical parameter values, a genetic algorithm (GA) was used and implemented in MATLAB (MathWorks, Natick, MA, USA). The GA improved the accuracy of the biogas yield by optimizing ADM1. According to the sensitivity analysis results, the disintegration constant (kdis = 0.191 d−1) and the hydrolysis rate constant for carbohydrates (khyd_ch = 1.624 d−1) are critical parameters for optimising biogas production from maize silage. The study revealed that the mathematical model, through optimization, enhances the efficiency of the biogas digester.
The optimization of biogas potential using a kinetic model, response surface methodology (RSM) and instrumental evidence for the biodegradation of tannery fleshing during anaerobic digestion was studied by Mahendiran et al. (2023) [130]. A key aspect of the study methodology is the enzymatic pretreatment of tannery fleshing using trypsin and papain. Pretreatment with the trypsin enzyme is applicable to the leather industry, food processing, and bioremediation, whereas papain is obtained from papaya fruit. The enzyme accelerates the anaerobic digestion process. The study, conducted on a batch scale with a retention time of 45 days, produced daily biogas, measured using the water displacement method. For the kinetic analysis, the authors employed the first-order model and the modified Gompertz model to optimize the biogas production, using the following parameters (temperature, pH, and hydraulic retention time) as independent variables. The first-order and modified Gompertz model equations proposed in the study are presented in Equations (10) and (11), respectively, using ORIGIN 2019 software.
where P the cumulative biogas yield P0 is the ultimate biogas yield, k and t represents the first-order constant and time, respectively.
where L is the lag phase duration, R is the biogas production rate, and P0 is the biogas potential at time (t). According to the kinetic model, the authors reported that both the first-order and the Gompertz models provided a good fit, with R2 ranging from 0.994 to 0.998 and 0.983 to 0.993, and RMSE ranging from 1.74 to 2.80 and 1.04 to 3.43, respectively. This signifies a satisfactory fit between the model and data. Regarding daily biogas production, the first-order kinetic model yielded 402.27–439.82 mL/gVS, whereas the modified Gompertz model reported 407.70–446.44 mL/gVS.
Mohammadianroshanfekr et al. (2024) [27] published an article on the kinetic modelling and optimisation of biogas production from food waste and cow manure co-digestion in a batch digester of six 1000 mL Erlenmeyer flasks. The study aimed to determine the best model for predicting biogas potential, considering first-order kinetics (Equation (12)), the transference model (Equation (13)), the modified Gompertz model (Equation (14)), and the modified logistic model (Equation (15)). These model equations were used to analyse the experimental results. The choice of food waste and cow manure was attributed to their high availability and large quantities in which they are generated. Specifically, food waste provides a consistent and rich source of organic matter, whereas cow manure is available from agricultural activities for biogas production. Equations (12)–(15) shows the model’s equation used in the study.
In the above equations, y is the cumulative biogas yield (mL/g VS), P refers to the ultimate biogas yield (mL/g VS), K is the first-order kinetic rate constant (1/day), Rm is known as the maximum biogas production (mL/g VS/day), whereas λ is the lag phase time, and e is the mathematical constant (Neper number). The study focused on determining the coefficient of determination (R2) and the root mean square error (RMSE) to assess the goodness-of-fit of each model. Table 3 presents the study’s findings.
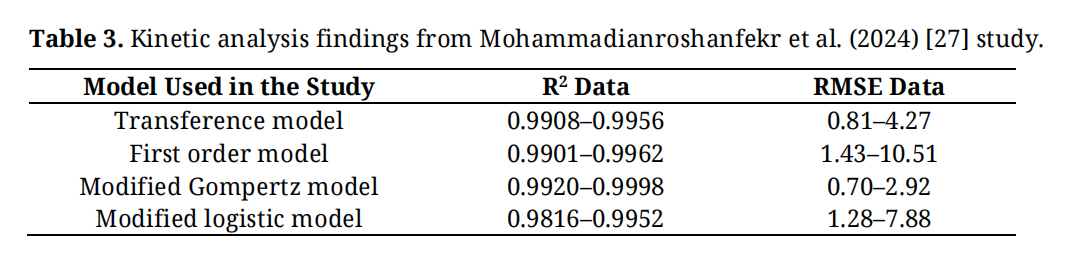 Table 3. Kinetic analysis findings from Mohammadianroshanfekr et al. (2024) [27] study.
Table 3. Kinetic analysis findings from Mohammadianroshanfekr et al. (2024) [27] study.
From Table 3, it is evident that all the models used in the study exhibited high predictive accuracy, with the modified Gompertz model providing the best fit, as indicated by an R2 value of 0.999. The finding indicates that the modified Gompertz model is reliable and has potential for predicting biogas yield, thereby optimising anaerobic digestion processes.
Prediction of biogas yieldTo investigate the performance of three kinetic models for the prediction of biomethane yield of combined oxidation and nanoparticle additive pretreated Xyris capensis, Olatunji et al. (2021) [131] carried out the study. The kinetic model used in the study includes first-order, logistic function and modified Gompertz models. In their study, the predicted biomethane yields from different kinetic models under varying treatment conditions were compared with the experimental yield. First, the Xyris capensis was pre-treated using an oxidative technique with a prepared Piranha solution. The choice of this pretreatment method is based on its economic viability and ease of application. During the pre-treatment, the sample was dried in an oven at 60 °C for approximately 6 h, until the desired moisture content was achieved. For characterization and experimentation, the sample was kept in a zip-lock plastic bag. The software used for data analysis and graphing in the study was OriginPro 2024, and ANOVA was used to determine significance. The Equations (16)–(18) were employed in the study as a first-order, logistic function and a modified Gompertz model.
where T is the total biomethane released during the retention time (mL/g VSadded), To is the specific biomethane potential of the feedstock (mL/g VSadded), r is the biomethane release rate constant (mL/day), and t is the time (days).
where B is the total biomethane yield at time (mL/g VSadded), H is the biomethane strength of the feedstock (mL/g VSadded), μm is the optimum biomethane released rate, λ is the lag phase time (days), and t is the digestion time (days).
where G is the cumulative biomethane during the digestion time (mL/g VSadded), A is the biomethane potential of the feedstock (mL/g VSadded), μm is the highest biomethane produced, λ is the lag phase time (days), and t is the digestion time (days). However, the findings from the study show that the three models used are characterised by acceptable values for λ, R² (0.8269–0.9978), RMSE (3.0193–156.3094), Akaike information criterion (AIC) of 65.6708–175.1098, and percentage difference (0.0329–4.7751). This further revealed that the first-order kinetic model is more accurate for predicting the combined pretreated substrate, whereas the modified Gompertz model was the most accurate for simulating the kinetics of the pretreated substrate's biomethane yield.
As in the previous study, Roberts et al. (2023) [81] employed five kinetic models to predict methane yields from oranges and bananas. The models employed in the study include the Gompertz, logistic, first-order, Richard, and transfect models. The model parameters used in the study are the maximum methane potential yield (A), the lag phase, the biogas rate constant (k), the shape factor coefficient (d), the incubation time (t), and the biogas production rate (μm). It is worth noting that the input data for the various models were obtained from experimental data. The study was programmed and analysed in MATLAB using input parameters. Regarding methane yield, all five models reached a maximum of 352.9 mL, with the Richard and the transfect model in 15 days (the fastest). Using banana and orange as substrates, the study reported comparative findings. For banana as the substrate, a phase lag (days) of 0.317 was reported for the five models used in the study, excluding the first-order model. The methane potential and production for the Gompertz model were 352.9 and 2.05 mL, respectively. For the orange as the substrate, 0.31775 was reported for the five models, excluding the first-order term. The methane potential and production for the Gompertz model were 464.6 and 3.332 mL, respectively. Considering the shape factor coefficient, this was reported for the Richard model only as 0.334 and 1.67 for banana and orange, respectively. The first-order kinetic model was reported to exhibit the greatest deviation, due to the absence of a lag phase and the methane production rate. The Gompertz model had the least deviation from the actual yield, with a deviation of 113.2%, followed by the logistic, Richards, first-order, and the transfect models, with deviations of 187.4%, 218.0%, 250.0%, and 803.0%, respectively. Therefore, in general, all models showed methane yield prediction accuracies of over 99.23% for banana peel, with the Gompertz model being the most accurate (76% cumulative deviation) and the transfer model the least accurate (274.7% cumulative deviation). In contrast, for orange peels, the methane yield prediction was 99.49%. The most accurate model was the Gompertz and first-order kinetic model, with a cumulative deviation of 20.67%. The least accurate model was the transfer model, with 112 cumulative deviations. This study clearly confirms that kinetic modelling can predict biogas production regardless of the parameters and conditions used.
Validation of experimental dataWith reference to studies by Gaur & Suthar (2017) [132], Pererva et al. (2020) [119], and Tahir Iqbal et al. (2021) [133], kinetic modelling is a method used to validate experimental data. This indicates that its application involves comparing experimental results with predictions derived from the kinetic model. In doing so, it ensures accuracy and prediction performance under various conditions. To achieve this, models such as the first-order, Monod, or Gompertz models are applied to experimental data, with the aim of evaluating the goodness of fit using statistical methods (R2 and RMSE). A few studies on the validation of experimental data via kinetic modelling are briefly discussed.
The kinetic modelling of thermophilic anaerobic digestion of Lemnaceae for biogas production was researched by Lower et al. (2025) [120]. The study used four kinetic models, which were fitted to the experimental data. These models include the first-order, modified Gompertz, transference, and logistic functions, as shown in Equations (21)–(24).
Equation (21) is also known as the developed conventional method for microbial growth kinetics. Specifically for the prediction of maximum methane potential (mL CH4 g−1 VS). In Equation (19), M(t) is the cumulative methane, Mu is the ultimate methane yield, and k is the hydrolysis constant.
where Rm is the maximum methane rate of production (mL g−1 VS day−1), λ and is the lag phase length (day). Equation (20) is used to estimate biochemical methane production and microbial growth-dependent processes (such as anaerobic digestion) [134]. For experimental data, the modified Gompertz model, as shown in Equation (20), has been shown to fit well to data from mesophilic anaerobic digestion of selected wastes (food waste, food waste co-digested with duckweed, and waste-activated sludge co-digested with duckweed) [135–137].
where M(t) is the cumulative methane, Mu is the ultimate methane yield, t is the digestion time (days), Rm is the maximum methane rate of production and λthe lag phase length (days). According to Ugwu & Enweremadu (2019) [138] and Fernández-Rodríguez et al. (2023) [139], the transference model in Equation (21) can be used to determine the same parameters as the modified Gompertz model (Equation (20)), as it depends on the microbial growth rate. In terms of experimental data, the transference model supports the best fit for thermophilic anaerobic data, primarily from waste-activated sludge and fat, oil, and grease [140]. The same is evident in the study by Lower et al. (2025) [120], which used Lemnaceae as a feedstock for biogas production. Other studies showing the transfer function model's outperformance in thermophilic data for biogas production include Pagliaccia et al. (2016) [141], Donoso-Bravo et al. (2010) [43], and Li et al. (2015) [87]. However, the logistic function model is presented in Equation (22), applicable in predicting the same parameter as the transference and modified Gompertz equation that governs microbial growth kinetics [120]
For the logistic function shown in Equation (22), it is assumed that methane production is proportional to the volume of accumulated biogas. Hence, this behaviour slows down as it approaches the maximum methane yield, as reported by Wang and Guo (2024) [134]. Based on experimental data and validation, the logistic function model is reported to have outperformed the first-order model, particularly for mesophilic anaerobic digestion of cow dung, and to have provided accurate predictions that align with the experimental data. In Equation (22), all the parameters have the same definition as previously used. Nevertheless, the authors reported that all four models used in the study accurately estimate methane production from the biomass. This was based on the difference between the predicted and observed methane production, which is reported to be a very small range of 0.03%–5.43%. Furthermore, the first-order model provided the best fit to the experimental data (R2 < 0.99), whereas the transfer model reported R2 > 0.98. Overall, the first-order and transfer models predicted the closest maximum methane production in Lemnaceae.
In a similar study, Ugwu & Enweremadu (2019) [138] focused on the biodegradability and kinetic studies on biomethane production from okra (Abelmoschus) waste. The study was motivated by the lack of research on biodegradability and degradation dynamics in biomethane production, as noted in the literature. First, the proximate and ultimate analyses of the okra samples were carried out, including moisture content, total solids, volatile solids, pH, C/N ratio, and ash content (%), among others. As part of the study's methodology, the authors employed nonlinear regression analysis using the curve-fitting tool in MATLAB 2015b to evaluate the growth function. Equations (23)–(27) represent the various kinetic models used in the study. These are first-order, Stannard, transference, logistic function and modified Gompertz models.
In defining the parameters, B is the cumulative specific methane production (mL/gVS); B0 is the maximum specific methane production potential (mL/gVS); Rmax is the maximum specific rate of methane production (mL/gVS-d), Exp = e = 2.718282, λ is the lag phase in days, k is the methane production per constant (day), and t is the digestion time (days) as well as p is the slope of growth. In the study, Table 4 presents the R2 and RMSE for the employed models.
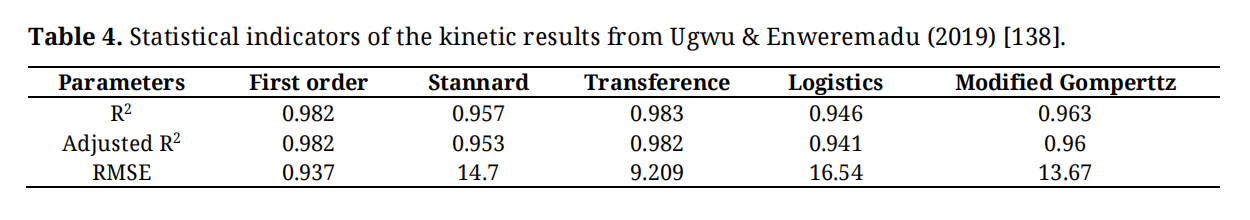 Table 4. Statistical indicators of the kinetic results from Ugwu & Enweremadu (2019) [138].
Table 4. Statistical indicators of the kinetic results from Ugwu & Enweremadu (2019) [138].
In Table 4, the statistical indicators of the four models employed in the study, in terms of R², Adj R², and RMSE, range from 0.946 to 0.983, 0.941 to 0.982, and 9.209 to 13.67, respectively. Based on Table 1, the first-order and transfer model, in terms of RMSE, reported values < 10, indicating good model predictions [142]. This indicates that both models are the best at predicting biochemical methane production. However, in response to the methane yield, the experimental and theoretical methane yields were 270.98 and 444.48 mL/gVS, respectively. Meanwhile, the model simulation yielded values ranging from 267.5 mL/g VS to 270.89 mL/g VS.
Scaling up processesThe accurate description of how models developed at a laboratory or small scale can predict the behaviour of a system at a larger or industrial scale is referred to as scaling-up via kinetic modelling. Scaling processes aim to identify the fundamental physical and chemical mechanisms that tend to be invariant across different scales, which relate to system performance in terms of size change [143].
Abilmazhinov et al. (2023) [114] developed a new mathematical model of anaerobic fermentation under industrial-scale operation. The authors aimed at creating an intelligent system for predicting biogas yield in a bioenergy unit, thereby determining and evaluating its energy capacity. The study utilised data from the biogas station in Kurma village, the Republic of Kazakhstan. In terms of the mathematical modelling, 20 to 26 different parameters were employed for industrial and practical applications. These parameters include micro- and macronutrients, which are lacking in previous models in the literature. Equation (28) presents the proposed linear regression model, which is recognized for its effectiveness and wide application in industrial biogas production.
In this case, y is the dependent variable (to be estimated), x is the explanatory variable, a is the regression coefficient, and n is the total number of observations. The study's findings showed that the biogas output has an average approximate error of 4.49 and an R2 value of 0.96. This indicates that the model was accurate (with a percentage error of less than 5%) and of higher quality (with an R-squared value closer to 1). The overall analysis of the model shows a good fit to the data. In terms of scaling the model, this tends to accommodate different changing parameters, according to the study. Hence, the proposed model was said to differ from the similar ADM1 in terms of accuracy, scalability, and flexibility. Further analysis to determine the dependence between biogas yield and the different parameters used in the study reveals R2 values of 0.6125, 0.6451, 0.6217, and 0.8229 for biogas yield, carbon emissions, nitrogen emissions, and volatile acid amount, respectively.
The mathematical model of a laboratory-scale plant for the bio-digestion of slaughterhouse effluents for biogas production was published by Martinez et al. (2012) [142]. The purpose of this study is to develop a simplified model that fits data obtained from a laboratory-scale prototype digester, thereby providing a reliable description of the dynamic process. In terms of feeding mode, the system is a continuous digester with a volume of 6.2 L, fed with a mixture of waste from the municipal slaughterhouse. The author reported that the parameters used in the study were fitted in two steps–first, adjusting parameters for the measured variables, such as COD, acetic and propionic acids, and methane, using a genetic algorithm. The remaining are fitted to values obtained from the literature. According to the model proposed by Aceves-Lara et al. (2005) [144], which is employed in the study, it consists of six equations (Equations (29)–(34)). The first equation addresses the control of substrate evolution, while the other three focus on the evolution of the bacterial population. Thus, the seventh equation (Equation (35)) presented by the authors describes the production of methane from acetic acid.
In Equations (29)–(35), X1 is the population of the bacterial acidogenesis in relation to the COD degradation. The X2 is the methanogenic bacteria, while the X3 represents the acetogenic bacteria. Additionally, the COD is represented by S1, S2, and S3, which denote the concentrations of acetic and propionic acid, respectively. It is interesting to note that the kinetic model related to scaling processes, as described by the COD in the equation, is based on the Monod kinetic equation, whereas the Haldane kinetic expression is applicable to the degradation of propionic and acetic acids. The study found that the model provides a reliable prediction of the system variables used (COD, propionic and acetic acids, and methane). Hence, it has proven to be the best model for adding an original component to the equation describing the process, especially for the anaerobic digestion of co-digestion of slaughterhouse effluent. Based on the digester's scaling process relative to kinetic modelling, the model can be used to test different controllers to efficiently achieve both waste degradation and methane generation.
Previous Studies on the Kinetic Modelling of Biogas ProductionTian et al. [145] employed anaerobic co-digestion (AcoD) experiments, varying mixing ratios, total solid concentrations, and inoculum amounts using pig manure and rice straw. The study employed a 1:5 mixing ratio of pig manure to rice straw, with a total solid content of 12%. With 15% inoculum, this yielded the highest biogas production of 553.79 mL/g VS. The optimal combination enhanced the cooperative index (CPI > 1), thereby indicating a synergistic effect that improved methane production and system stability. Based on the kinetic modelling result, which was analyzed using MATLAB, the modified Gompertz model's fitting results showed high agreement with experimental data, validating its accuracy in simulating methanogenic processes. The value of R2 for the kinetic parameters used in the study includes 0.997–0.999 for mixing ratio, 0.997–0.999 for the TS content, as well as 0.9096–0.999 for the inoculum amount. Additionally, the phase lags (days) for mixing ratio are 2.96–6.83, 3.88–5.86, and 3.33–6.16 for TS content, mixing ratio, and inoculum amount, respectively.
The Anaerobic Digestion Model No. 1 (ADM1) is a widely used mathematical model that simulates anaerobic digestion processes, predicting methane production and process outcomes [146]. The model is said to have been developed by the International Water Association (IWA) Task Group for Mathematical Modelling of Anaerobic Digestion Processes, specifically the ADM1 model. This incorporates biochemical reactions, microbial populations and physicochemical processes. The ADM1 involve implementing a set of differential equations to describe substrate degradation, biomass growth and biogas production. ADM1 has been shown to accurately predict methane production and process dynamics; however, it requires careful calibration and validation to ensure accuracy. When using the ADM1, it is recommended to carefully calibrate the model using experimental data to ensure accurate predictions. Additionally, sensitivity analysis is necessary to identify critical parameters and understand the impact on model outcomes. For model validation, it is necessary to utilise independent datasets to ensure the model’s accuracy and reliability. Finally, it is recommended to extend or modify the model as needed to accommodate specific applications or substrates, while maintaining a common platform. Therefore, following these recommendations, researchers can effectively utilize ADM1 to optimise anaerobic digestion processes, predict methane production and improve process efficiency.
A biochemical methane potential (BMP) study utilized the modified Gompertz model to simulate methane production from co-digesting different substrates [147]. The study used goat manure (GM) and cotton gin trash (CGT) as substrates for anaerobic digestion to produce biomethane. Part of the study's methodology involved fitting the modified Gompertz equation to experimental BMP data to estimate key kinetic parameters, including R2, RMSE, and lag phase time. The findings reported a lag phase of 0.5 ± 0.1–4.8 ± 0.7 days, R2 of 0.983 ± 0.003–0.993 ± 0.002, and RMSE of 0.502–1.373. The modified Gompertz model accurately predicted methane production and provided valuable insights into the synergistic effects of co-digestion on biogas yield. Further, the modified Gompertz model predicted, and the experimental biomethane value was similar. The highest synergistic effect index of 15.6 ± 4.7 was observed in GM: CGT; 30:70 co-digestion. Additionally, the study revealed that the model optimized co-substrate ratios, predicted methane production, and identified optimal conditions for enhanced biogas production. The application of the aforementioned model in the study helps improve the efficiency and stability of anaerobic co-digestion systems, ultimately enhancing biogas production and reducing greenhouse gas emissions.
The kinetic simulation for thermophilic biogas production, using data from market wastes co-digested with rumen matter, was conducted by Mbugua et al. (2023) [108]. The market waste consists of mixed fruit and vegetable waste co-digested with slaughterhouse waste at 55 °C and a pH of 6.8–7.2. As part of the study's methodology, the data were fitted into linear, exponential, Gaussian, logistic, and modified Gompertz kinetic models. The Equations (36)–(40) present the kinetic models used in the study.
where y is the biogas rate of production in ml/gm/day; t is the digestion time (days); a and b are known as the intercept and slope of the graph, respectively. In Equation (37), it is expected that biogas production will increase linearly with time and, at the maximum point, decrease linearly to zero as time increases further. For the exponential kinetic model;
where c is a constant (d−1), and other parameters or variables remain the same as previously defined. The exponential kinetic model, as shown in Equation (38), indicates that the rate of biogas production increases exponentially with time and then decreases to zero exponentially after reaching a certain point. To simulate the rate of biogas production, the Gaussian kinetic model, as shown in Equation (38), describes the process and follows a normal distribution across HRT.
In this case, t0 is the time required for the maximum biogas production to occur. Equation (40) represents the logistic function kinetic model.
where y is the cumulative production of biogas (ml/gm); k is the kinetic constant rate (1/day); t is the HRT, and a and b are constants. Finally, the modified Gompertz model presented in Equation (40) is a statistical regression model used to illustrate batch biogas digestion. The equation shows that methane production depends on bacterial growth in a logarithmic relationship [148].
where Y is the cumulative specific production of biogas (ml/gm); A is the biogas production potential (mL/gm), U is the maximum biogas production (mL/gm/day), and λ is the lag phase period (days). The kinetic variables of the models reported in the study are presented in Table 5.
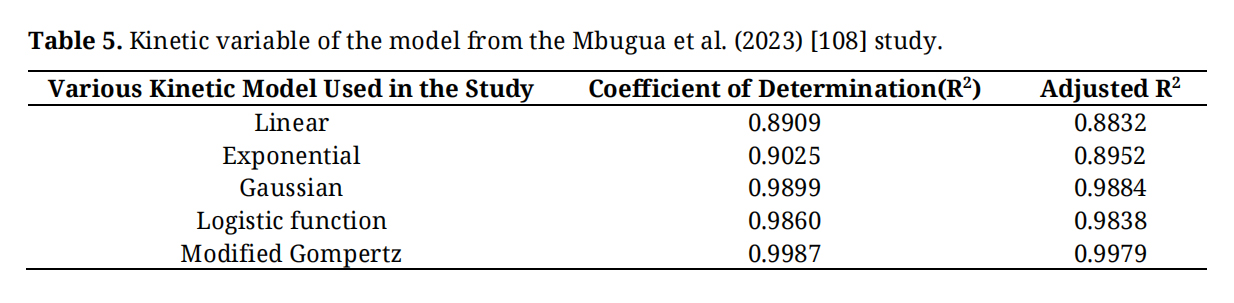 Table 5. Kinetic variable of the model from the Mbugua et al. (2023) [108] study.
Table 5. Kinetic variable of the model from the Mbugua et al. (2023) [108] study.
Similar to the previous study by the same author, Mbugua et al. (2024) [149] conducted a kinetic modelling study of biogas production from market waste inoculated with Abattoir waste. Blended twenty market wastes were inoculated with rumen fluid at mesophilic temperature for a seven-day hydraulic retention time. The results from the study were used for the kinetic studies by fitting them to linear, exponential, Gaussian, and Modified Gompertz kinetic models, as shown in Equations (41)–(44).
where B1 is the biogas production rate (L kg−1 d−1) at time t (day) over the digestion period, a1 is the intercept (L kg−1 d−1), and b1 is the slope (L kg−1 d−1). From Equation (1), it follows that biogas generation depends on HRT.
where C is a constant (d−1) and other parameters remain the same as previously defined.
where t0 is the time (day) at which the biogas peak occurred. In Equation (39), the biogas production rate and its microbial kinetic growth are functions of the normal distribution, as determined by the breakdown period.
From the above equations, Table 6 presents the findings of the kinetic evaluation of biogas production, as determined by the coefficient of determination (R2).
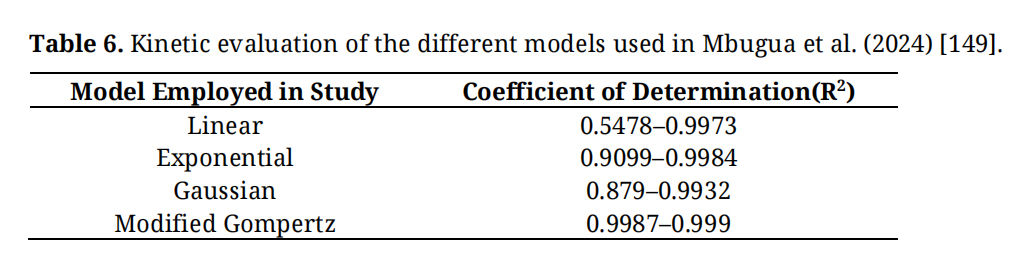 Table 6. Kinetic evaluation of the different models used in Mbugua et al. (2024) [149].
Table 6. Kinetic evaluation of the different models used in Mbugua et al. (2024) [149].
Evidently, from Table 6, the Modified Gompertz model yielded the highest accuracy, ranging from 0.9987 to 0.999. However, the biogas production from the models is reported to have a high accuracy of 25.96 mL/g, compared to the experimental result of 23.58 mL/g. Mathematically, this shows a deviation of 2.87%.
The study of the kinetics of biogas production from anaerobic digestion of vinasse waste was conducted by Usmani et al. (2021) [150]. Considering the waste generated, vinasse is recognized as a valuable sugar ethanol residue suitable for biogas production due to its high COD content. In addition to the kinetic study, the study aimed to enhance biogas production using vinasse wastewater as a substrate and Citadel BioCat + as a source of methanogenic bacteria. The authors designed a batch biogas digester for the experiment, employing online measurement of the methane rate. Equation (46) presents the first-order kinetic model for the biodegradability of organic wastewater.
The differential rate of Equation (45) can be expressed as
Here, CA,t and CA,i represent the concentration of COD at the beginning and subsequent times, respectively. Using the modified Gompertz model (Equation (49)) and nonlinear regression, the production of biogas is said to correspond to the specific growth of methanogenic bacteria in the anaerobic digester.
p is the cumulative biogas production (mL/g-COD), A is the production of biogas potential (mL/g-COD), U is the highest biogas production rate (mL/g-COD/d) and λ is the minimum biogas production time, t is the HRT, and e is exp (2.7183). Table 6 presents the findings from the study by Usmani et al. (2021) [150].
From Table 7, it is evident that temperature and the presence of a biocatalyst significantly affect biogas production. Increasing the temperature (37 °C), without a catalyst, enhances the production of biogas and reduces the time for anaerobic digestion.
To thoroughly investigate the performance of organic degradation and identify the interaction between the reduction of organic components and methane yield, Li et al. (2018) [40] laid the groundwork for this study. The study focused on kinetic studies of organic degradation and its impact on enhancing methane production from 12 different types of food waste. Five models were used in the study, as shown in Equations (50)–(54): exponential kinetics, the Fitzhugh model, the cone model, the transference function, and the modified Gompertz model.
Similarly to previous equations where applicable, B is the cumulative organics reduction for a digestion time (%), B0 is the ultimate organic reduction, k is the first order reduction rate (d−1), μm is the maximum organic reduction, λ is the lag time (days), and t and e are the time (days) and base of natural logarithm (2.7183), respectively. The authors calculated the parameter values for the kinetic model, as shown in Equations (51)–(55). With the use exponential model. It was reported that cumulative methane production increased exponentially from 385 to 627 mL/g volatile solids, with removal efficiencies of volatile solids, lipids, and proteins. On the other hand, the model simulation results revealed that higher methane production (530–548 mL/g VS), removal efficiencies of volatile solids (65–67%) and lipids (77.8–78.2%), and 54.7–58.2% protein were achieved with a short digestion retention time. This is true for the carbohydrate content of 54.7–58.2%, which is said to be higher than 47.6% and 28.3% for protein and lipids, respectively.
Sensitivity Analysis in Relation to Kinetic Modelling of Anaerobic Digestion for Biogas ProductionSensitivity analysis arises from the non-linear dynamics of anaerobic digestion, the uncertain feedstock quality, and the sensitivity of environmental conditions, all of which are due to the complexity of interacting biochemical processes. Based on this, anaerobic digestion is considered highly susceptible to instability. To address these problems, researchers have conducted sensitivity analyses of kinetic models for biogas production via anaerobic digestion. This is essential, as it is a critical component in the kinetic modelling of biogas production, where input parameters are identified. Furthermore, the sensitivity analysis is important for identifying how model outcomes differ as assumptions are altered. According to Boutoute et al. (2021) [151], sensitivity analysis is helpful for verifying the validity of model assumptions and identifying parameters. To demonstrate a stronger correlation with experimental data and to calibrate the model parameter, sensitivity analysis is employed. Some of these parameters include kinetic rate constants, stoichiometric coefficients, and mass transfer coefficients. The input parameter is said to affect the model output. A scoping review conducted by Barahmand and Samarakoon (2022) [152] on the application of sensitivity analysis in anaerobic digestion modelling revealed that ADM1 is the most commonly used model for sensitivity analysis. For instance, this can be evident in a study by Bułkowska et al. (2025) [129], in which the results of the sensitivity analysis underscore the importance of adjusting ADM1 parameters to reflect substrate-specific properties. The study noted that the high sensitivity of methane production to the disintegration constant and the carbohydrate hydrolysis rate constant is consistent with the results of Guo et al. [153]. This implies that adjusting the stoichiometric and kinetic parameters for a specific fraction improves the fit of the ADM1 model for complex substrates such as sewage sludge. In the study by Boutoute et al. (2021) [151], a sensitivity analysis was developed to highlight the key parameters of a dry anaerobic digestion reactor model. The authors employed the defining screening design (DSD) and the multiple linear regression (MLR) analysis as the methodology for the sensitivity analysis. Both methods are said to require a small number of runs and are applicable only to the simplified anaerobic digestion model. Based on the sensitivity analysis results, it was found that the slow hydrolysis rate and high pH inhibition limit hydrolytic biomass production, consistent with the observed biogas flow rate. The mass transfer coefficient, determined in the sensitivity analysis, is reported to have a significant impact on pH via CO2 desorption. The authors recommend DSS and MLR as easy to apply to any type of model, given their simplicity and reliability. In another study, it was noted that simulation modelling of microbial proliferation has increased in recent years, aided by Monod and other advanced kinetic models. Based on that, Abubakar et al. (2022) [154] conducted a study on the Monod equation focusing on the sensitivity analysis of the kinetic growth in the model data. The authors examined the behaviour of the Monod parameter across different substrates (wastewater, animal residues, and lignocellulosic material) and biomass levels, using predictive data derived from empirical observations. The study's findings show that predicted data are more convenient and better capture the expected dynamics of the process than ideal experimental results. A sensitivity analysis was conducted in a study by Bekrit and Ogwu (2025) [100] to evaluate the influence of key kinetic and stoichiometric parameters on methane production. The analysis was employed using the one-at-a-time (OAT) method. Parameters used from the ADM1 include hydrolysis rate, decay coefficient, microbial yield and substrate uptake characteristics. It was reported that these parameters were perturbed individually within a range of ±10, while the others remained constant. The results of the finding revealed that the hydrolysis rate of carbohydrate exhibited the highest influence on methane output, followed by that of protein and lipids. As in the previous study, a sensitivity analysis was used to identify and evaluate the most sensitive kinetic parameter in a study by Zhao et al. (2019) [117]. The study reported that the hydrolysis rate constant of carbohydrate affected the biogas production the most.
Trade-Offs in Kinetic Modeling SelectionThe selection of the model mostly involved balancing bias and variance. The objective of bias-variance trade-offs is to find a model with the right balance of bias and variance to ensure a good generalisation to unseen data. Bias arises from an overly simplistic model that fails to capture the underlying patterns in the data, whereas variance arises from a complex model that treats noise as a pattern. This is the main essence of trade-offs. Trade-offs balance the model complexities with generalisation performance. To capture patterns in data, both simple and complex models are employed. A simple model has high bias and low variance, leading to underfitting, whereas a complex model has low bias and high variance, leading to the risk of overfitting [155]. However, empirical evidence shows that multiple models often achieve competitive results and exhibit similar overall performance, which could lead to different operational outcomes. This development leads to an implicit trade-off in model performance, thereby creating space to resolve and requiring new model selection tools. To this end, He and Shaposhnik [156] explore methods for comparing predictive models in an interpretable manner to uncover trade-offs. Based on that, the study proposed various models to synthesise ideas from supervised and unsupervised learning, dimensionality reduction, and visualisation. It was noted that, with the Python interface, researchers can better understand the broader impact of model selection choices. Evaluation, prioritisation, and selection of parameter requirements are said to be of tremendous importance and to affect the software development used in the model. This was the case in a study of trade-off analysis for requirement selection by Ruhe et al. (2003) [157]. The authors proposed a method, Quantitative WinWin, as a trade-off analysis tool to support requirement negotiation. The tool is beneficial in four ways. First, it uses the Analytical Hierarchy Process (AHP) to balance preferences across different classes of requirements. Secondly, it helps in requirement selection by predicting and rebalancing effort, time, and quality in the dealing model kinetics. This uses a simulation model prototype called GENSIM. Thirdly, the trade-off tool provides a decision-making solution that is based on a threshold for the degree of importance to find the best fit to the constraints. Fourthly, determine the non-dominated extension of the maximum achievable value under resource and quality constraints. Ruhe et al. (2003) [157] recommend the Quantitative Win-Win as a tool for selecting the most appropriate solution and for making actual kinetic modelling decisions. In a similar study, Bai et al. (2017) [158] developed an optimal trade-off model for portfolio selection with parameter sensitivity estimated from historical data. The model, a quadratic programming problem, was developed for three functions: risk measurement, maximum sensitivity, and minimum sensitivity. The study found that the model based on the optimal trade-off delivers stable returns with lower risk during the stress test.
Limitations of the StudyDifferent feedstocks exhibit unique characteristics, such as varying organic matter content, nutrient profiles, and inhibitory compounds, which can affect the accuracy and applicability of kinetic models. Based on model assumptions, it is noteworthy that kinetic models rely on simplifying assumptions, such as uniform microbial populations and ideal mixing conditions, which may not accurately represent real-world anaerobic digestion systems. Accurate estimation of kinetic parameters requires high-quality experimental data, which can be challenging to obtain, particularly for complex feedstocks or co-digestion systems. During the review, findings may not directly translate to industrial-scale anaerobic digestion systems, where factors such as temperature fluctuations, pH variations, and equipment design can impact process performance. Furthermore, the study did not account for all potential inhibitory factors, such as toxic compounds, heavy metals or excessive ammonia production, which can impact microbial activity and process stability. Having established this, the choice of kinetic model can significantly affect prediction accuracy, and different models may be more suitable for specific feedstocks or process conditions. Finally, the quality and reliability of the experimental data used for model calibration and validation can significantly impact the accuracy of kinetic model predictions, which is also a limitation of the study. Addressing these limitations and challenges, kinetic modelling becomes an even more effective tool for optimizing anaerobic digestion systems and promoting the widespread adoption of biogas production technologies.
Future Research OutlookThe authors outline possible areas that require further research in relation to the study as follows:
●
●
●
●
●
The study successfully reviewed the kinetic modelling of anaerobic digestion for biogas production from different feedstocks/wastes, which is regarded as a valuable tool for the understanding of process dynamics and optimization of methane yields. The literature establishes that the main and widely used kinetic models in the study include the modified Gompertz, Logistic, Transference, and First-order models. This helps researchers and operators gain insights into key parameters, including methane production rates, lag phase durations, substrate degradation kinetics, and the coefficients of determination, root mean square errors, and hydraulic retention times. Additionally, the knowledge gained from the kinetic models enables the design and operation of more efficient anaerobic digestion systems, tailored to specific feedstocks and process conditions. According to the review, these kinetic models have found their way in the anaerobic digestion of biogas production in terms of optimization of biogas digester, predicting the biogas yield, validation of experimental data and scaling processes. The review indicates that the modified Gompertz and logistic function models provide a better fit for mesophilic anaerobic digestion. This is due to the mesophilic process, which possesses a pronounced lag phase, an exponential production phase, and a stabilisation phase of bacterial growth, which acclimates the microbial community to the substrate at a lower temperature. Also, both kinetic models are known for predicting optimum biogas potential and production rate. Meanwhile, the transference kinetic model also provides a better fit for thermophilic anaerobic digestion. This is attributed to the fast digestion of thermophiles, which results in shorter or negligible lag phase and a quicker initial surge in gas production. As seen in the review, high rates of solids thermophilic digestion, such as FOG sludge, show a higher correlation coefficient. The main highlight of the review is the importance of selecting appropriate kinetic models and understanding their limitations to accurately predict biogas production and process outcomes. By doing so, it enhances biogas yields, reduces process inhibition and improves the overall efficiency and stability of anaerobic digestion systems. Furthermore, it can also facilitate the development of more effective strategies for optimizing co-digestion, substrate pre-treatment and process monitoring.
During the writing of this manuscript, no data were generated from the review.
NKS, KO and PM designed and conceptualized the study, analyzed the data. NKS and KO wrote the original and draft papers, and PM supervised the study. All authors have agreed to the publication of this manuscript.
The authors declare that they have no conflicts of interest.
This research received no external funding.
We would like to express our sincere appreciation to the Department of Science and Innovation (DSI), the Technology Innovation Agency (TIA), the National Research Foundation (NRF), and the Research Niche Area: Renewable Energy (Wind) at the University of Fort Hare, South Africa, for their support.
1.
2.
3.
4.
5.
6.
7.
8.
9.
10.
11.
12.
13.
14.
15.
16.
17.
18.
19.
20.
21.
22.
23.
24.
25.
26.
27.
28.
29.
30.
31.
32.
33.
34.
35.
36.
37.
38.
39.
40.
41.
42.
43.
44.
45.
46.
47.
48.
49.
50.
51.
52.
53.
54.
55.
56.
57.
58.
59.
60.
61.
62.
63.
64.
65.
66.
67.
68.
69.
70.
71.
72.
73.
74.
75.
76.
77.
78.
79.
80.
81.
82.
83.
84.
85.
86.
87.
88.
89.
90.
91.
92.
93.
94.
95.
96.
97.
98.
99.
100.
101.
102.
103.
104.
105.
106.
107.
108.
109.
110.
111.
112.
113.
114.
115.
116.
117.
118.
119.
120.
121.
122.
123.
124.
125.
126.
127.
128.
129.
130.
131.
132.
133.
134.
135.
136.
137.
138.
139.
140.
141.
142.
143.
144.
145.
146.
147.
148.
149.
150.
151.
152.
153.
154.
155.
156.
157.
158.
Sheron NK, Obileke K, Mukumba P. Kinetic Modelling of Anaerobic Digestion of Biogas Production from Different Wastes: A Review. J Sustain Res. 2026;8(2):e260044. https://doi.org/10.20900/jsr20260044.

Copyright © Hapres Co., Ltd. Privacy Policy | Terms and Conditions